Please consider submitting issues to GitHub which will enable tracking bug reports and suggestions. These will be closed and updated automatically when they are fixed – including notifications.
I believe this to be a bug with Avogadro:
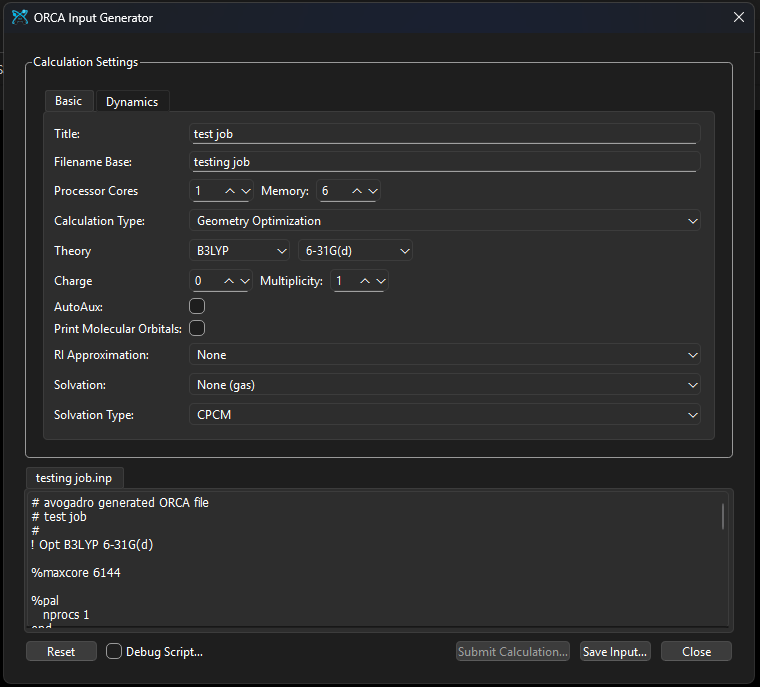
The ORCA input generator is non-functional across both the latest stable and continuous development builds on Windows. In the stable build, the “Submit Calculation” and “Input Generator” options remain grayed out despite the orca.py script being present in the default inputGenerators directory. In the continuous build, the menu options are active, but clicking them fails to launch the dialog box entirely.
Environment Information
Avogadro version: 1.103.0
Operating system and version: Windows 11 Pro, 25H2, 26200.7840
Expected Behavior
Upon loading a molecule (e.g., Caffeine), the Input→ORCA→ Submit Calculation options should be clickable.
Actual Behavior
“Submit Calculation” button is disabled (grayed out).
Steps to Reproduce
Install Avogadro 2 → Load Caffeine molecule → Go to Input>ORCA → Submit Calculation grayed out.
Please upload files if appropriate here (or via file-sharing service like Dropbox or Pastebin)
This is a future feature that’s not implemented yet.
Yes, you can’t submit a calculation at the moment. That’s part of the integration with the Molequeue app which handles calculations. You can save input and submit yourself.
If you think we should hide the button instead of having it visible but disabled, sure we could do that.
But you can absolutely generate ORCA input, which is of course the point of the input generator.
I am able to use the “Submit calculation” feature on Linux (version 1.103), and I actually find this an extremly useful feature, e.g., to set up a computational lab for students (so, please do not hide it!). As I read in other posts, Molequeue needs to be installed, set up and running in order to enable the “Submit calculation” button. I’ve tried with Orca (whose outut file is supported by Avogadro) and Psi4. The submission works fine, and the job is properly passed from Avogadro to Molequeue and run till sucess.
I find one possible bug, though. I marked the option “Open output when finished” and Avogadro seems to get the signals from Molequeue when the job is done, as it changes from “Waiting for job…” to “Fetching completed job information” when the job is completed. However, it never gets the info and indicates: “Avogadro timed out waiting for the finished job details from MoleQueue”. Instead, if I try to open the file from the folder where the job is done, the file “job.out” is properly loaded (which is a good way to bypass the issue). Actually, opening the job folder might be a better default behavior after finishing, as that would allow to select the file (out, fchk, allxyz…)
Glad to know it’s getting some use! Yes, it’s a nice system when everything is set up. IMHO, the biggest step will be to update MoleQueue so it’s a bit easier to get going (e.g., providing some defaults for running locally and some mechanisms to share default setups for use on clusters).
I’m pretty sure the most recent releases of Avogadro have had issues with receiving information.
It’s definitely fixed because the same protocol is used for automation (e.g., open a set of files, save screenshots, render orbitals).
Maybe I have something set up wrongly. Which file name is Avogadro expecting? I am asuming “job.out” (with Orca), is it right?
These are the messages I get in the terminal where I submit the job fom Avogadro:
When the job is submitted:
qt.core.qobject.connect: QObject::connect: No such signal Avogadro::MoleQueue::Client::errorReceived(int, uint, QString) in /home/javier/dev/Repos/Avogadro2/openchemistry/avogadrolibs/avogadro/molequeue/molequeuewidget.cpp:273
qt.core.qobject.connect: QObject::connect: (receiver name: 'widget')
qt.core.qobject.connect: QObject::disconnect: No such signal Avogadro::MoleQueue::Client::errorReceived(int, uint, QString) in /home/javier/dev/Repos/Avogadro2/openchemistry/avogadrolibs/avogadro/molequeue/molequeuewidget.cpp:278
qt.core.qobject.connect: QObject::disconnect: (receiver name: 'widget')
and when MoleQueue is finised:
qt.core.qobject.connect: QObject::connect: No such signal Avogadro::MoleQueue::MoleQueueWidget::jobUpdated(MoleQueue::JobObject) in /home/javier/dev/Repos/Avogadro2/openchemistry/avogadrolibs/avogadro/molequeue/molequeuedialog.cpp:110
qt.core.qobject.connect: QObject::connect: (sender name: 'widget')
qt.core.qobject.connect: QObject::connect: (receiver name: 'widget')
qt.core.qobject.connect: QObject::connect: No such signal Avogadro::MoleQueue::MoleQueueWidget::jobUpdated(MoleQueue::JobObject)
qt.core.qobject.connect: QObject::connect: (sender name: 'widget')
No - my comment is that in recent versions of Avogadro using Qt6, the “receive from Molequeue” is broken. It should be fixed now, but I haven’t had a chance to test it.
It doesn’t “expect” a filename. MoleQueue sends the file using an RPC message in JSON.
It’s on the TODO to test before 2.0 is released.