Several people have asked about a plot for relative energies, etc.
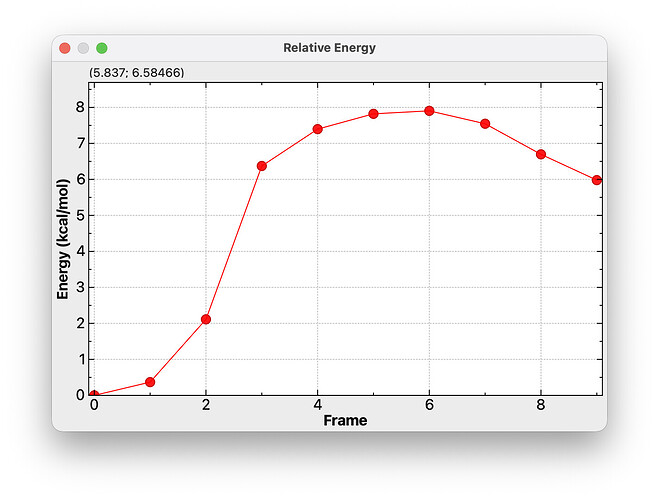
Here’s an example for an NEB trajectory.
My idea is that this “Plot Conformers…” dialog would allow you to switch between RMSD or relative energies. It would also offer a “Scale Energies” to convert from Hartree, eV, etc. to kcal/mol as needed. (In this case, the relative energies were in Hartree.)
And you can click on the plot to switch conformers.
Other ideas or suggestions?