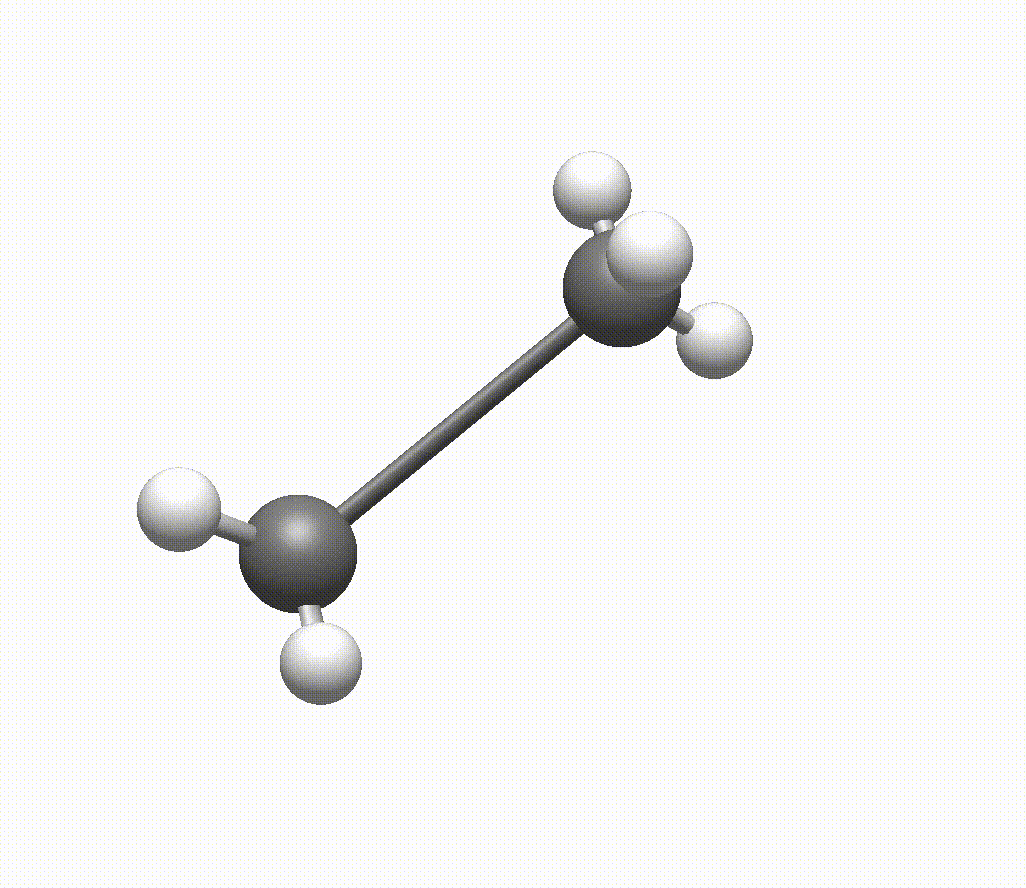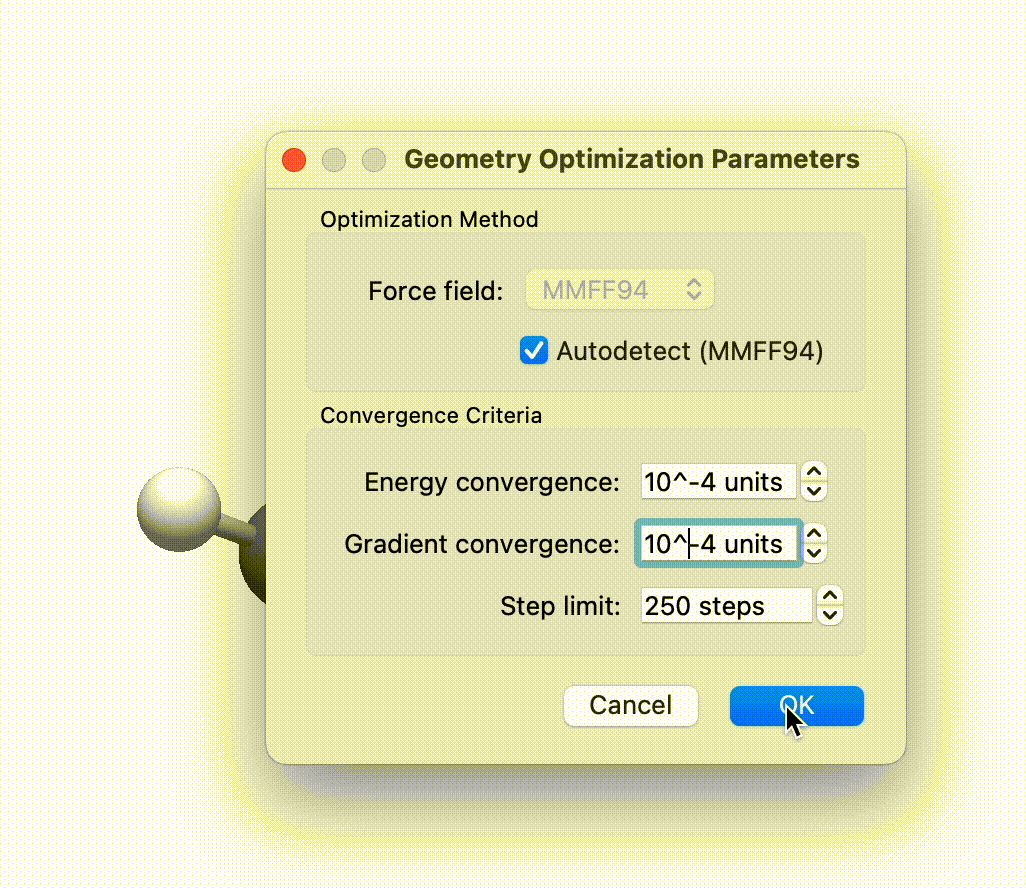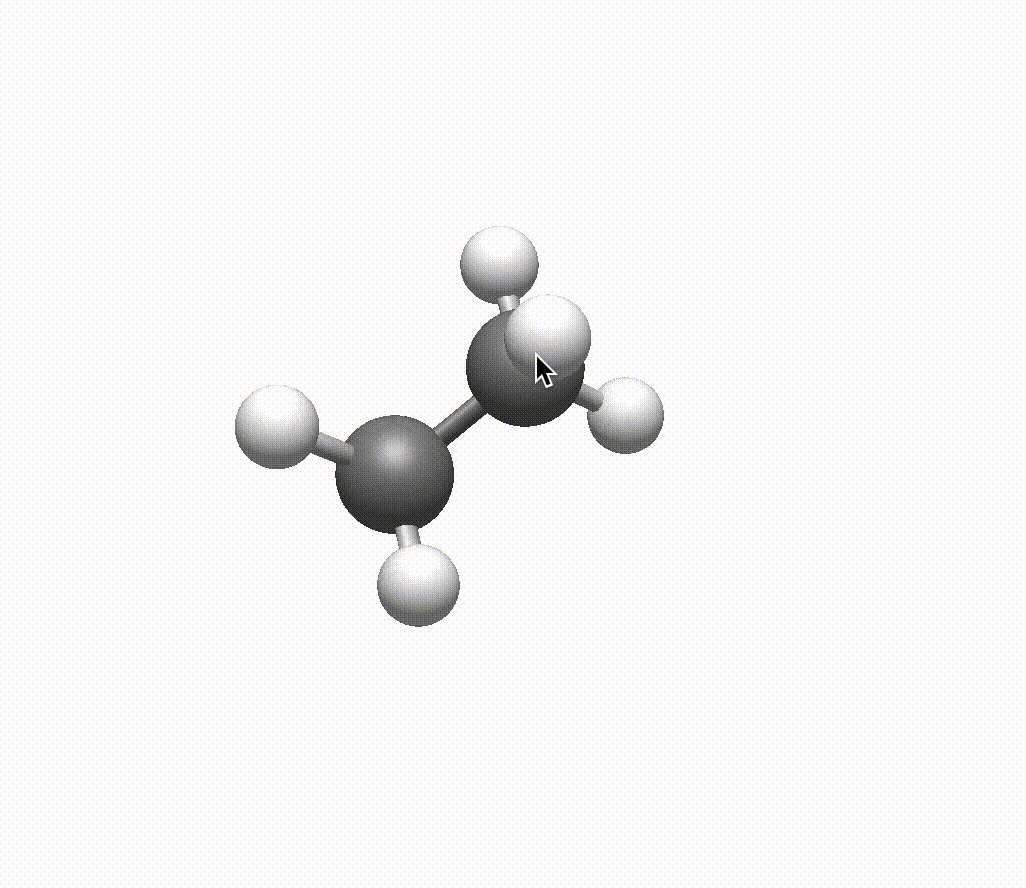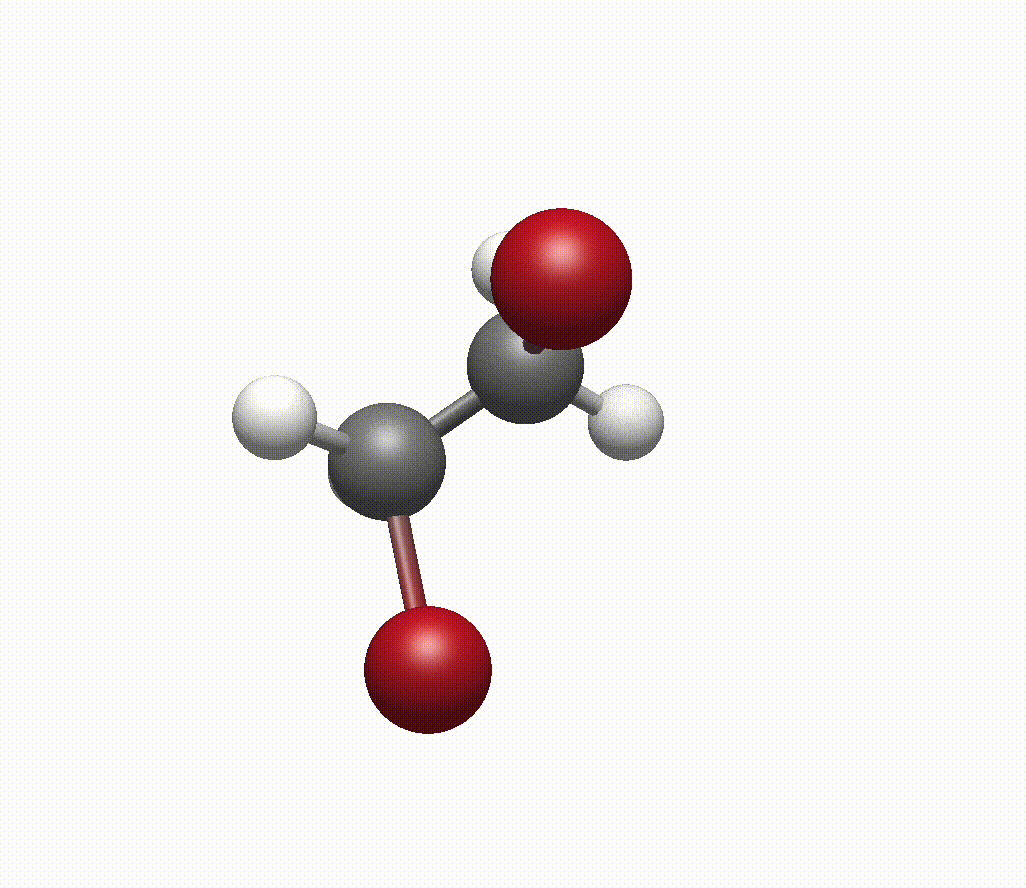I will shortly be merging in the new force field framework.
- You will be able to create and install Python scripts for optimizing molecules & unit cells
- e.g. ML potentials like ANI-1ccx and ANI-2x
- calculators for materials like ASE, OpenKIM, etc.
xtb-python like GFN-FF
- Each script can declare what it supports:
- unit cells or only molecules
- charged species
- radicals
- specific elements (e.g., only HCON or Ag / Au alloys, or C nanomaterials, etc.)
- analytical gradients
- Scripts get a copy of the current molecule in a range of file formats (e.g., a temporary file)
- Scripts return energies and/or gradients
Avogadro handles frozen atoms / constraints by modifying the forces / gradients back from the script. Right now, there’s just freeze / unfreeze selected atoms.
Still to come:
- Auto-optimize tool interface with “live” optimization. Real Soon Now™
- Full replacement for Open Babel optimization (e.g., right now, this is an extra option - 1.99 / 2.0 will merge them)
- Distance / angle / torsion constraints (e.g., debugging this took a bunch of time, so I haven’t had time to code these up)
Great work. Been waiting for this for a while, can’t wait to try it out!
Please let us know how it goes - it’s obviously really new so I’m sure there will be bugs.
Would you give a bit more info about how to interface with this framework? Trying to figure out how to play with something a little more powerful than LJ when testing.
Best,
Sam
If you conda install -c conda-forge openbabel you’ll get the MMFF94 and UFF force fields. If you conda install -c xtb-python you should get GFN-FF.
I was working on an Open Babel interface that doesn’t require the Python bindings, but had a bunch of hard to debug issues. I’m making progress on that and will likely implement UFF internally as well.
Thanks i’ll give it a shot!
Ok - Installed the conda packages. How do I get avogadro to ‘read’ them? I’m not seeing any options other than LJ. Do I need to develop my own script following something like:
https://two.avogadro.cc/develop/scripts/energy.html
Thanks!
No, a bunch of scripts should be installed with 1.98.
What OS are you using? Have you restarted Avo2 after installing the conda packages?
If you launch from the command-line prompt, you should currently see messages about registering the force fields.
Hmm. Ok, I think I see whats going on. I think it isn’t finding my python interpreter, ubuntu jammy Jellyfish.
Ok, I set my python path and it found the force fields. Thanks!
Yes, we need some better tools for finding Python and selecting Python environments.