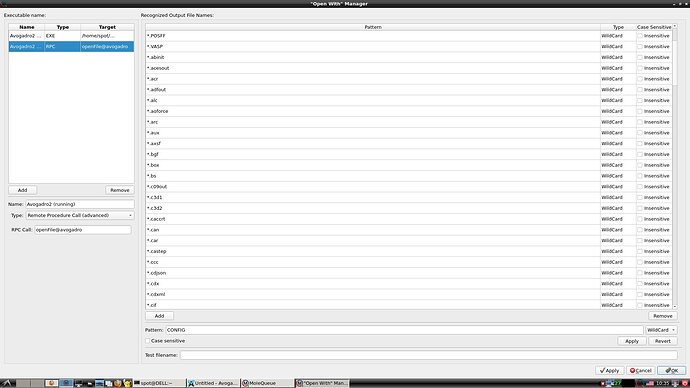
I believe this to be a bug with Avogadro2
Environment Information
Avogadro version: 2
Operating system and version: Arch Linux 4.19.88-1-lts
Expected Behavior: Trying to Open finished job file into RUNNING Avogadro2
Actual Behavior: Avogadro2 opened empty (untitled) molecule instead of completed job file with optimized geometry.
Steps to Reproduce: Quantum → Input Generator job was submitted for nwchem calculations from within avogadro2. Completed nwchem job cannot be automatically/manually open into RUNNING avogadro2 , but only into new avogadro2 instance - which worked just fine.
Molqueue log shows NO entries.
Popup massage says “Avogadro timed out waiting for the finished job details from MoleQueue.” and then “The job did not complete successfully.”, which is wrong - job was successfully completed.
I am running avogadro2 from terminal which shows the following upon job submission to nwchem calculations:
QObject::connect: No such signal MoleQueue::Client::errorReceived(int, uint, QString) in /home/spot/openchemistry/avogadrolibs/avogadro/molequeue/molequeuewidget.cpp:288
QObject::connect: (receiver name: ‘widget’)
QObject::disconnect: No such signal MoleQueue::Client::errorReceived(int, uint, QString) in /home/spot/openchemistry/avogadrolibs/avogadro/molequeue/molequeuewidget.cpp:293
QObject::disconnect: (receiver name: ‘widget’)
QObject::connect: No such signal Avogadro::MoleQueue::MoleQueueWidget::jobUpdated(MoleQueue::JobObject) in /home/spot/openchemistry/avogadrolibs/avogadro/molequeue/molequeuedialog.cpp:125
QObject::connect: (sender name: ‘widget’)
QObject::connect: (receiver name: ‘widget’)
QObject::connect: No such signal Avogadro::MoleQueue::MoleQueueWidget::jobUpdated(MoleQueue::JobObject)
QObject::connect: (sender name: ‘widget’)
What version (e.g. Git, 1.90, 1.91, etc.) are you using of Avogadro and Molequeue?
I’m also going to ping @mhanwell since the Molequeue integration is on his side. I’d guess some of the signals are out of sync between Molequeue and that version of Avogadro.
Avogadro 1.91.0
Avogadrolibs 1.92.0
MoleQueue 09.0-4-g0511c9a
These were current on Git today when I cloned them.
Thanks, I’ll take a look… I’ll be busy the next couple of days, but please post this at https://github.com/openchemistry/avogadrolibs/issues and I’ll definitely make sure it gets fixed for the next release.
I will try to take a better look at this, I have been a little neglectful of the MoleQueue piece as we have worked on Avogadro improvements and other web-based features. Let me see what I can see.
As a follow up to my post I am adding the output of make during avagodro2 compiling.
Apparently, when I choose in ccmake option BUILD_AVOGADRO_CLIENT_SERVER ON
the following error will follow:
““CMake Error at ~/openchemistry/openchemistry-build/prefix/lib/cmake/vtk-8.2/vtkModuleAPI.cmake:140 (message):
Requested modules not available:
vtkParallelCore
Call Stack (most recent call first):
~/openchemistry/openchemistry-build/prefix/lib/cmake/vtk-8.2/VTKConfig.cmake:143 (vtk_module_config)
avogadro/qtplugins/clientserver/CMakeLists.txt:7 (find_package)””
How to let make know locations of vtkParallelCore?
My current setup:
Avogadro version: 1.93.0
Avogadrolibs: 1.93.0
Qt: 5.14.2
MoleQueue 0.9.0-4-g0511c9a
OS: Arch Linux
Version 5.4.24-1-lts
Compiler gcc version 9.3.0
I don’t think you need BUILD_AVOGADRO_CLIENT_SERVER to do what you want, but I’d leave that to @mhanwell
I also think the VTK version is beyond 8.2 - my build has VTK 9.0. I’d recommend making sure you have the latest version of the git master for openchemistry and avogadrolibs