Hey all, after dealing with problems with the download plugins menu not loading anything (fixed that by installing OpenSSL) I am now trying to load an ORCA 5.0.4 .out file and I want to view the molecular orbitals. From some previous posts, I heard it was possible by loading the files through cclib, but I have no idea how to do that. I initially tried using the download plugins feature to download cclib, but after a quick restart of Avogadro, no new plugins appeared in the extensions menu, and using the open command in the file menu yielded no molecular orbitals. For some background information, I am running Windows 11, grabbed the nightly build today on 11/30/23 at 9:45pm, I have python 3.12 installed, and have checked that it is added to the system path. I have no clue where to go from here.
If your Orca file has MO / basis information, you don’t need cclib with 1.98.1 – it has a native Orca reader. (And if the file doesn’t have the info, cclib won’t be able to help either.)
If you want to use cclib, there’s no extra menu command. It’s just an option to help read files. Just make sure you’ve installed the Python package too (e.g., pip3 install cclib)
As an example:
benzene.out.gz (119.8 KB)
After opening a file from an ORCA 5.0.4 output, I use the Analysis menu and create surfaces, but when I pick, for example, the HOMO and try to generate the surface, Avogadro crashes. If I recall correctly from Avogadro 1, there was just a menu to pick the orbital to display and it automatically brought it up. Is there something like that in Avogadro 2 that I am just missing or is it actually the create surfaces menu and there is some bug?
https://www.dropbox.com/scl/fo/0y6kz8l3rbxv0sev6ru6r/h?rlkey=3jbn6b71vuqc4mm62jmho2xqd&dl=0
First off, there’s not yet the “orbital table” from Avogadro 1.
Beyond that, it’s definitely a bug - I was able to reproduce your crash. Not sure why, but I’ll look into it this evening.
Interesting. I also tried a different output file of a metal complex I am studying, and it was able to generate the MOs, though I couldn’t visualize them for some reason.
Make sure the “Meshes” display type is checked.
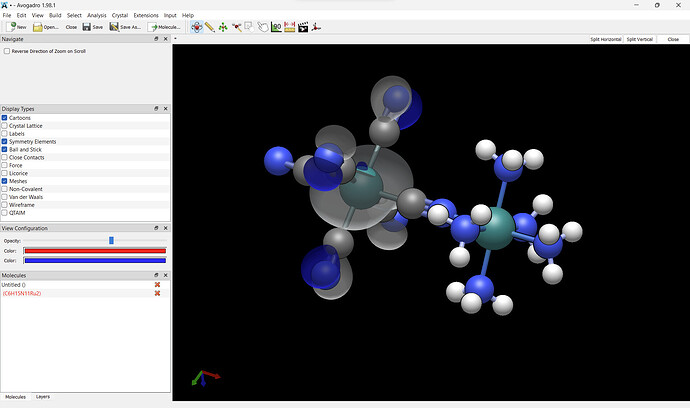
Lovely. Although I did notice that the coloration of the orbitals isn’t quite correct, it should be red and blue but instead gray and blue, is this an error on my end?
Edit: After trying to calculate one more orbital and display it, avogadro 2 crashed.
I’d suggest changing the opacity first, then clicking on the red to change the color. Certainly if you want it red / blue, it shoudl be red / blue. ![]()
If you’re willing to share other Orca files that crash, please send them my way.
If you need a workaround, you can always use orca_2mkl and generate a Molden file. The Orca import code is new and probably has more bugs than the Molden reader.
Looks like part of the issue is that you’re using “g” basis functions (e.g., def2-QZVP) which aren’t implemented yet. On the plus side, I have an old pull request that was waiting for some test files. ![]()
I have a bunch of files using QZVP, some are just some pentanediones with only carbon hydrogen and oxygen, and some are with halogens up to iodine. I have in total about 20 files, some are just optimizations, some are just frequencies, some are both. Any particular requests or should I just dump them all in a dropbox?
Thanks, I think for now, I’m good - let me make sure I can get what you’ve sent working. At that point, it’s worth testing more options.