Hi,
How can i build epsilon-polylysin (ε-Polylysine) instead normal a-polylysin?
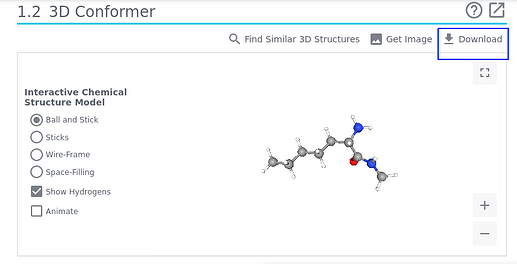
There is entry 20719176 of PubChem, an open access database provided by the National Institute of Health/NIH. Go to section 1.2
and click in the top right corner (marked above): one option allows you to save a .sdf file locally you eventually may read e.g., with Avogadro.
With a running instance of Avogadro, choose Ctrl + O (or in the GUI of Avogadro, File → Open) to read this file just saved. Avogadro will ask you how to process this input; in the intermediate interface, choose Avogadro: MDL as file reader, and confirm by Enter. On occasion, you have to tap once (left mouse button) on Avogadro’s canvas to complete the display of the newly accessed model.
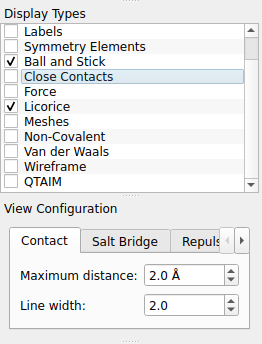
If not already selected (typically left hand side of the GUI), you may choose the representation of a ball-stick models:
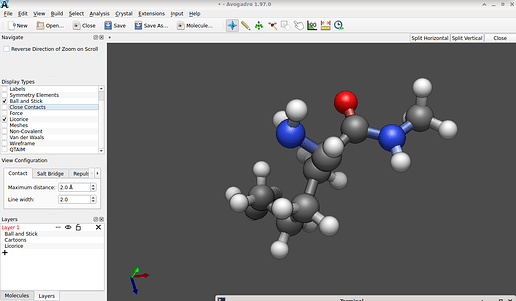
(In case you got a model which still is flat-2D, you have to run a quick, computationally affordable geometry optimization. There are two way to launch this: either by pressing Ctrl + Alt + O simultaneously as a shortcut, or – using the GUI – by mouse clicking on Extensions → OpenBabel → Optimize Geometry to yield one plausible conformer. Usually, the geometry assigned by NIH however is retained, and this extra step is not necessary.) And this should offer you the model of a monomer of ε-polylysine:
edit: correction of the import of 2D/3D data.
It shouldn’t be a “flat” 2D geometry. That implies you downloaded the 2D file, not “Conformer3D_CID_20719176.sdf” which should certainly be a 3D file. (I just checked)
Note in the URL, there’s a record_type=3d
In any case, to build up a polymer, you can, for the moment, copy-and-paste. Certainly it would be nice to have some better polymer and oligopeptide builders for Avo2, and there are some design plans, but as of yet no one to work on them.
This topic was automatically closed 3 days after the last reply. New replies are no longer allowed.