I probably won’t have a chance to get all those features in Mercury anytime soon - they’re great and I’ve definitely used them, especially for generating molecular clusters.
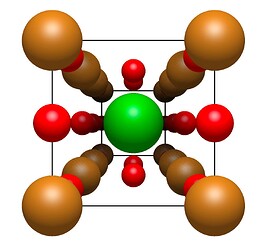
What I did manage, is generating all the copies on the unit cell boundaries, e.g. KBr
I need to do a bit more (e.g., automating bond perception) but it seems like it’s a step in the right direction for preview images.
What do you think the default should be when loading inorganic .cif files?
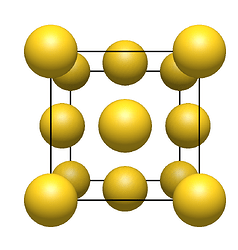
- symmetry unique atoms (e.g., one gold atom)
- four gold atoms
- capped unit cell