Hello, dear support,
PubChem database offers json-format files, like this one, Centromere protein V-like protein 1 (human) | Protein Target - PubChem ,
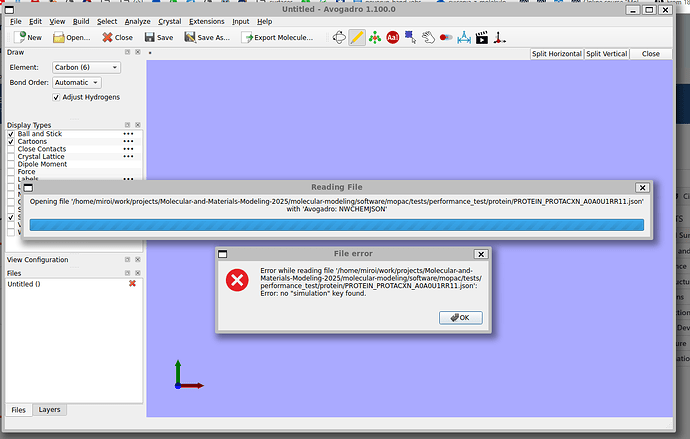
but Avogadro2 (nor JMol) can not read it. Avogadro2 can read SDF files, but they are not present at each PubChem compound.
Please help to find a workaround.
Open Babel has some support for JSON from PubChem compounds (e.g., as pcjson format) but this particular file has very little chemical information. Look at it yourself .. it looks more like a set of links than something with 3D molecular information.
"Record": {
"RecordType": "Protein",
"RecordAccession": "A0A0U1RR11",
"RecordTitle": "Centromere protein V-like protein 1 (human)",
"RecordExternalURL": "https://www.ncbi.nlm.nih.gov/protein/A0A0U1RR11",
"Section": [
{
"TOCHeading": "Protein Information",
"Description": "General information about a protein",
"DisplayControls": {
"HideThisSection": true
},
"Section": [
{
"TOCHeading": "Taxonomy",
"Description": "Source organism of this protein.",
Now if you dig through this carefully, there is a FASTA sequence. There’s also something that has a link to the AlphaFold DB but it seems like it would be easier for you to just download that PDB or mmCIF file.
Please - take a look at that particular JSON file yourself. Let me know how you think Avogadro should handle it. Besides “oh here’s a FASTA, I’ll generate some arbitrary non-folded protein” I’m not sure what the program should do.
Incidentally, if you have a good suggestion on searching the AlphaFold DB, I’d be very interested - seems like it would be a useful feature.
For example, it seems easy if you have the accession number:
https://alphafold.ebi.ac.uk/files/AF-A0A2K6V5L6-F1-model_v4.pdb
Or more specifically, this returns JSON:
https://alphafold.ebi.ac.uk/api/prediction/Q5VSL9 ⇒
https://alphafold.ebi.ac.uk/files/AF-Q5VSL9-F1-model_v4.pdb